Written by Godson Obielum✏️
In this tutorial, we’ll demonstrate how to build a YouTube video search application using Angular and RxJS. We’ll do this by building a single-page application that retrieves a list of videos from YouTube by passing in a search query and other parameters to the YouTube search API.
We’ll use the following tools to build our app.
- TypeScript, a typed superset of JavaScript that compiles to plain JavaScript and provides type capabilities to JavaScript code
- Angular, a JavaScript framework that allows you to create efficient and sophisticated single-page applications
- RxJS, a library for composing asynchronous and event-based programs by using observable sequences. Think of RxJS as Lodash but for events
You should have a basic understanding of how these tools work to follow along with this tutorial. We’ll walk through how to use these tools together to build a real-world application. As you go along, you’ll gain practical insight into the core concepts and features they provide.
You can find the final code in this GitHub repository.
Prerequisites
You’ll need to have certain libraries installed to build this project locally. Ensure you have the Node package installed.
We’ll use Angular CLI v6.0.0 to generate the project, so you should ideally have that version installed to avoid weird errors later on.
Project setup
1. Structure the application
Before we start writing code, let’s conceptualize the features to be implemented in the application and determine the necessary components we’ll need.
We’ll keep it as simple as possible. At the core, we’ll need to have an input element that allows the user to type in a search query. That value will get sent to a service that uses it to construct a URL and communicate with YouTube’s search API. If the call is successful, it will return a list of videos that we can then render on the page.
We can have three core components and one service: a component called search-input for the input element, a component called search-list for rendering the list of videos, and a parent component called search-container that renders both the search-input and search-list components.
Then we’ll have a service called search.service. You could think of a service as the data access layer (DAL), that’s where we’ll implement all the relevant functionality that’ll enable us to communicate with the YouTube search API and handle the subsequent response.
In summary, there will be three components:
search-containersearch-inputsearch-list
The search-input and search-list components will be stateless while search-container will be stateful. Stateless means the component never directly mutates state, while stateful means that it stores information in memory about the app state and has the ability to directly change/mutate it.
Our app will also include one service:
search.service
Now let’s dive into the technical aspects and set up the environment.
2. Set up the YouTube search API
We’ll need to get a list of YouTube videos based on whatever value is typed into the input element. Thankfully, YouTube provides a way that allows us to do exactly that by using the YouTube search API. To gain access to the API, you’ll need to register for an API token.
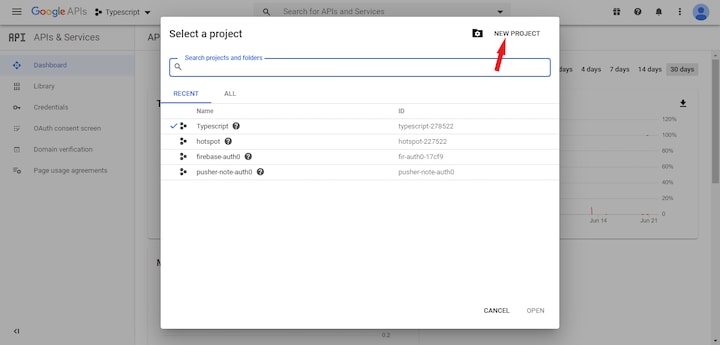
First, if you don’t already have one, you’ll need to sign up for a Google account. When that’s done, head over to the Google developer console to create a new project.
Once the project is successfully created, follow the steps below to get an API token.
- Navigate to the credentials page by clicking on
Credentialslocated on the sidebar menu - Click on the
+ CREATE CREDENTIALSbutton located at the top of the page and selectAPI key.A new API key should be created. Copy that key and store it somewhere safe (we’ll come back to it shortly) - Head over to the API and Services page by clicking on
APIs & Serviceslocated at the top of the sidebar - Click on
ENABLE APIs AND SERVICESat the top of the page. You’ll be redirected to a new page. Search for the YouTube Data API and click on theYoutube Data API v3option. Once again, you’ll be redirected to another page. ClickEnableto allow access to that API
With that done, we can start building out the application and the necessary components.
3. Scaffold the application
Create a directory for the application. From your terminal, head over to a suitable location on your system and issue in the following commands.
# generate a new Angular project
ng new youtube-search `
# move into it
cd youtube-search
This uses the Angular CLI to generate a new project called youtube-search. There’s no need to run npm install since it automatically installs all the necessary packages and sets up a reasonable structure.
Throughout this tutorial, we’ll use the Angular CLI to create our components, service, and all other necessary files.
Building the application
1. Set up the search service
Before we build the search service, let’s create the folder structure. We’ll set up a shared module that will contain all the necessary services, models, etc.
Make sure you’re in your project directory and navigate to the app folder by running the following command.
cd src/app
Create a new module called shared by running the following command in the terminal.
ng generate module shared
This should create a new folder called shared with a shared.module.ts file in it.
Now that we have our module set up, let’s create our service in the shared folder. Run the following command in the terminal.
ng generate service shared/services/search
This should create a search.service.ts file in the shared/services folder.
Paste the following code into the search.service.ts file. We’ll examine each chunk of code independently.
// search.service.ts
import { Injectable } from '@angular/core';
import { HttpClient } from '@angular/common/http';
import { map } from 'rxjs/operators';
import { Observable } from 'rxjs';
@Injectable({
providedIn: 'root'
})
export class SearchService {
private API_URL = 'https://www.googleapis.com/youtube/v3/search';
private API_TOKEN = 'YOUR_API_TOKEN';
constructor(private http: HttpClient) {}
getVideos(query: string): Observable <any> {
const url = `${this.API_URL}?q=${query}&key=${this.API_TOKEN}&part=snippet&type=video&maxResults=10`;
return this.http.get(url)
.pipe(
map((response: any) => response.items)
);
}
}
First, take a look at the chunk of code below.
import { Injectable } from '@angular/core';
import { HttpClient } from '@angular/common/http';
import { map } from 'rxjs/operators';
import { Observable } from 'rxjs';
@Injectable({
providedIn: 'root'
})
[...]
In the first part of the code, we simply import the necessary files that’ll help us build our service. map is an RxJS operator that is used to modify the response received from the API call. HttpClient provides the necessary HTTP methods.
@Injectable() is a decorator provided by Angular that marks the class located directly below it as a service that can be injected. { providedIn: 'root'} signifies that the service is provided in the root component of the Angular app, which in this case is the app component.
Let’s look at the next chunk:
[...]
export class SearchService {
private API_URL = 'https://www.googleapis.com/youtube/v3/search';
private API_TOKEN = 'YOUR_API_KEY';
constructor(private http: HttpClient) {}
getVideos(query: string): Observable <any> {
const url = `${this.API_URL}?q=${query}&key=${this.API_KEY}&part=snippet&type=video&maxResults=10`;
return this.http.get(url)
.pipe(
map((response: any) => response.items)
);
}
}
We have two private variables here. Replace the value of API_KEY with the API token you got when you created a new credential.
Finally, the getVideos method receives a search query string passed in from the input component, which we’ve yet to create. It then uses the http get method to send off a request to the URL constructed. It returns a response that we handle with the map operator. The list of YouTube video details is expected to be located in the response.items object and, since we’re just interested in that, we can choose to return it and discard the other parts.
Due to the fact that the search service uses the HTTP client, we have to import the HTTP module into the root component where the service is provided. Head over to the app.module.ts file located in the app folder and paste in the following code.
import { BrowserModule } from '@angular/platform-browser';
import { NgModule } from '@angular/core';
import { AppComponent } from './app.component';
import { HttpClientModule } from '@angular/common/http';
@NgModule({
declarations: [
AppComponent
],
imports: [
HttpClientModule,
BrowserModule
],
providers: [],
bootstrap: [AppComponent]
})
export class AppModule { }
That’s basically all for the search service. We’ll make use of it soon.
2. Add a video interface file
Let’s quickly set up an interface file. A TypeScript interface allows us to define the syntax to which any entity must adhere. In this case, we want to define certain properties each video object retrieved from the Youtube search API should contain. We’ll create this file in the models folder under the shared module.
Run the following command in your terminal.
ng generate interface shared/models/search interface
This should create a search.interface.ts file. Copy the following code and paste it in there.
export interface Video {
videoId: string;
videoUrl: string;
channelId: string;
channelUrl: string;
channelTitle: string;
title: string;
publishedAt: Date;
description: string;
thumbnail: string;
}
Interfaces are one of the many features provided by TypeScript. If you aren’t familiar with how interfaces work, head to the TypeScript docs.
Setting up the stylesheet
We’ll be using Semantic-UI to provide styling to our application so let’s quickly add that.
Head over to the src folder of the project, check for the index.html file, and paste the following code within the head tag.
<link rel="stylesheet" type="text/css" href="https://cdn.jsdelivr.net/npm/fomantic-ui@2.8.5/dist/semantic.min.css">
Your index.html file should look something like this:
<!doctype html>
<html lang="en">
<head>
<meta charset="utf-8">
<title>YoutubeSearch</title>
<base href="/">
<meta name="viewport" content="width=device-width, initial-scale=1">
<!-- Added Semantic Ui stylesheet -->
<link rel="stylesheet" type="text/css" href="https://cdn.jsdelivr.net/npm/fomantic-ui@2.8.5/dist/semantic.min.css">
<link rel="icon" type="image/x-icon" href="favicon.ico">
</head>
<body>
<app-root></app-root>
</body>
</html>
Setting up the stateless components
1. Develop the search input component
The next step is to set up the stateless components. We’ll create the search-input component first. As previously stated, this component will contain everything that has to do with handling user input.
All stateless components will be in the components folder. Make sure you’re in the app directory in your terminal before running the following command.
ng generate component search/components/search-input
This creates a search-input component. The great thing about using Angular’s CLI to generate components is that it creates the necessary files and sets up all boilerplate code, which eases a lot of the stress involved in setting up.
Add the following HTML code to the search-input.html file. This is just basic HTML code and styling using semantic UI:
<div class="ui four column grid">
<div class="ten wide column centered">
<div class="ui fluid action input">
<input
#input
type="text"
placeholder="Search for a video...">
</div>
</div>
</div>
Take note of the #input line added to the input element. This is called a template reference variable because it provides a reference to the input element and allows us to access the element right from the component.
Before we start working on the component file, there are a few things to handle on the input side:
- Set up an event listener on the input element to monitor whatever the user types
- Make sure the value typed has a length greater than three characters
- It’s counterintuitive to respond to every keystroke, so we need to give the user enough time to type in their value before handling it (e.g., wait 500ms after the user stops typing before retrieving the value)
- Ensure the current value typed is different from the last valu. Otherwise, there’s no use in handling it
This is where RxJS comes into play. It provides methods called operators that help us implement these functionalities/use cases seamlessly.
Next, add the following code in the search-input.component.ts file.
// search-input.component.ts
import { Component, AfterViewInit, ViewChild, ElementRef, Output, EventEmitter } from '@angular/core';
import { fromEvent } from 'rxjs';
import { debounceTime, pluck, distinctUntilChanged, filter, map } from 'rxjs/operators';
@Component({
selector: 'app-search-input',
templateUrl: './search-input.component.html',
styleUrls: ['./search-input.component.css']
})
export class SearchInputComponent implements AfterViewInit {
@ViewChild('input') inputElement: ElementRef;
@Output() search: EventEmitter<string> = new EventEmitter<string>();
constructor() { }
ngAfterViewInit() {
fromEvent(this.inputElement.nativeElement, 'keyup')
.pipe(
debounceTime(500),
pluck('target', 'value'),
distinctUntilChanged(),
filter((value: string) => value.length > 3),
map((value) => value)
)
.subscribe(value => {
this.search.emit(value);
});
}
}
Let’s take a look at a few lines from the file above.
-
ViewChild('input')gives us access to the input element defined in the HTML file previously.'input'is a selector that refers to the#inputtemplate reference variable we previously added to the input element in the HTML file -
ngAfterViewInitis a lifecyle hook that is invoked after the view has been initialized. In here, we set up all code that deals with the input element. This ensures that the view has been initialized and we can access the input element, thereby avoiding any unnecessary errors later on
Now let’s look at the part of the code found in the ngAfterViewInit method.
- The
fromEventoperator is used to set up event listeners on a specific element. In this case, we’re interested in listening to thekeyupevent on the input element - The
debounceTime()operator helps us control the rate of user input. We can decide to only get the value after the user has stopped typing for a specific amount of time — in this case, 500ms - We use the
pluck('target','value')to get the value property from the input object. This is equivalent toinput.target.value -
distinctUntilChanged()ensures that the current value is different from the last value. Otherwise, it discards it. - We use the
filter()operator to check for and discard values that have fewer than three characters - The
mapoperator returns the value as anObservable. This allows us to subscribe to it, in which case the value can be sent over to the parent component (which we’ve yet to define) using theOutputevent emitter we defined.
That’s all for the search-input component. We saw a tiny glimpse of just how powerful RxJS can be in helping us implement certain functionalities.
2. Develop the search list component
Now it’s time to set up the search-list component. As a reminder, all this component does is receive a list of videos from the parent component and renders it in the view.
Because this is also a stateless component, we’ll create it in the same folder as the search-input component. From where we left off in the terminal, go ahead and run the following command.
ng generate component search/components/search-list
Then head over to the search-list.component.ts file created and paste the following code in there.
// search-list.component.ts
import { Component, OnInit, Input } from '@angular/core';
import { Video } from '../../../shared/models/search.interface';
@Component({
selector: 'app-search-list',
templateUrl: './search-list.component.html',
styleUrls: ['./search-list.component.css']
})
export class SearchListComponent implements OnInit {
@Input() videos: Video[];
constructor() { }
ngOnInit() {
}
}
The file above is fairly straightforward. All it does is receive and store an array of videos from the parent component.
Let’s take a look at the HTML code, switch to the search-input.html file, and paste in the following code.
<div class="ui four column grid">
<div class="column" *ngFor="let video of videos">
<div class="ui card">
<div class="image">
<img [src]="video.thumbnail">
</div>
<div class="content">
<a class="header" style="margin: 1em 0 1em 0;">{{ video.title }}</a>
<div class="meta">
<span class="date" style="font-weight: bolder;">
<a [href]="video.channelUrl" target="_blank">{{ video.channelTitle }}</a>
</span>
<span class="ui right floated date" style="font-weight: bolder;">{{ video.publishedAt | date:'mediumDate' }}</span>
</div>
<div class="description">
{{ video.description?.slice(0,50) }}...
</div>
</div>
<a [href]="video.videoUrl" target="_blank" class="extra content">
<button class="ui right floated tiny red right labeled icon button">
<i class="external alternate icon"></i>
Watch
</button>
</a>
</div>
</div>
</div>
In the file above, we simply loop through the array of videos in our component and render them individually, this is done using the *ngFor directive found in the line above:
<div class="column" *ngFor="let video of videos">
Building the stateful component
Let’s create the parent component, search-container. This component will directly communicate with the search service sending over the user input and then pass the response to the search-list component to render.
Since the search-container is a stateful component, we’ll create this in a different directory than the other two components.
In the terminal once again, you should still be in the app directory. Type in the following command.
ng generate component search/container/search-container
Before we start writing code, let’s take a step back and outline what we want to achieve. This component should be able to get user inputs from the search-input component. It should pass this over to the search service, which does the necessary operations and returns the expected result. The result should be sent over to the search-list component, where it will be rendered.
To implement these things, paste the following code into the search-container.component.ts file.
// search-container.component.ts
import { Component } from '@angular/core';
import { SearchService } from 'src/app/shared/services/search.service';
import { Video } from 'src/app/shared/models/search.interface';
@Component({
selector: 'app-search-container',
templateUrl: './search-container.component.html',
styleUrls: ['./search-container.component.css']
})
export class SearchContainerComponent {
inputTouched = false;
loading = false;
videos: Video[] = [];
constructor(private searchService: SearchService) { }
handleSearch(inputValue: string) {
this.loading = true;
this.searchService.getVideos(inputValue)
.subscribe((items: any) => {
this.videos = items.map(item => {
return {
title: item.snippet.title,
videoId: item.id.videoId,
videoUrl: `https://www.youtube.com/watch?v=${item.id.videoId}`,
channelId: item.snippet.channelId,
channelUrl: `https://www.youtube.com/channel/${item.snippet.channelId}`,
channelTitle: item.snippet.channelTitle,
description: item.snippet.description,
publishedAt: new Date(item.snippet.publishedAt),
thumbnail: item.snippet.thumbnails.high.url
};
});
this.inputTouched = true;
this.loading = false;
});
}
}
In the code above, the handleSearch method takes in the user input as an argument. It then communicates with the getVideos method in the search service passing in the input value as an argument.
The subscribe function invokes this service call and the response from the getVideos method is passed to it as the items argument. We can then filter out the necessary values needed and add that to the videos array in the component.
Let’s quickly work on the HTML, paste this into search-container.html and we’ll go through it after:
<div>
<app-search-input (search)="handleSearch($event)"></app-search-input>
<div *ngIf="inputTouched && !videos.length" class="ui four wide column centered grid" style="margin: 3rem;">
<div class="ui raised aligned segment red warning message">
<i class="warning icon"></i>
<span class="ui centered" style="margin: 0 auto;">No Video Found</span>
</div>
</div>
<div *ngIf="loading" style="margin: 3rem;">
<div class="ui active centered inline loader"></div>
</div>
<app-search-list *ngIf="!loading" [videos]="videos"></app-search-list>
</div>
In the file above, we simply render both child components, search-input and search-list, and add the necessary input binding to the search-list component. This is used to send the list of videos retrieved from the service to the component. We also listen to an event from the search-input component that triggers the handleSearch function defined earlier.
Edge cases are also handled, such as indicating when no videos are found, which we only want to do after the input element has been touched by the user. The loading variable is also used to signify to the user when there’s an API call going on.
By default in every Angular application, there’s a root component, usually called the app-root component. This is the component that gets bootstrapped into the browser. As a result, we want to add the search-container component to be rendered there. The search-container component renders all other components.
Open the app.component.html file and paste the code below.
<div class="ui centered grid" style="margin-top: 3rem;">
<div class="fourteen wide column">
<h1 class="ui centered aligned header">
<span style="vertical-align: middle;">Youtube Search </span>
<img src="/assets/yt.png" alt="">
</h1>
<app-search-container></app-search-container>
</div>
</div>
Testing out the Application
We’re all done! Now let’s go ahead and test our app.
In your terminal, run the following command to kickstart the application.
ng serve
You may encounter an error similar to ERROR in ../../node_modules/rxjs/internal/types.d.ts(81,44): error TS1005: ';' expected. This doesn’t have to do with the code but rather the RxJS package installation. Luckily, there’s a very straightforward and easy solution to that.
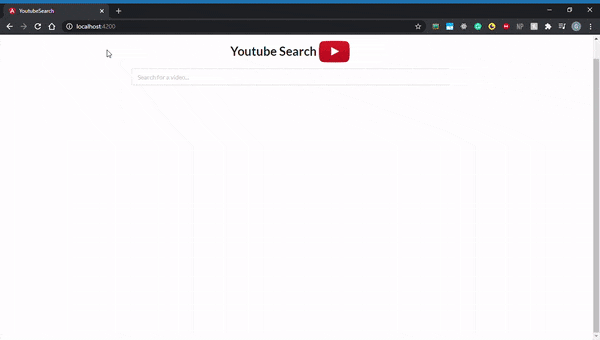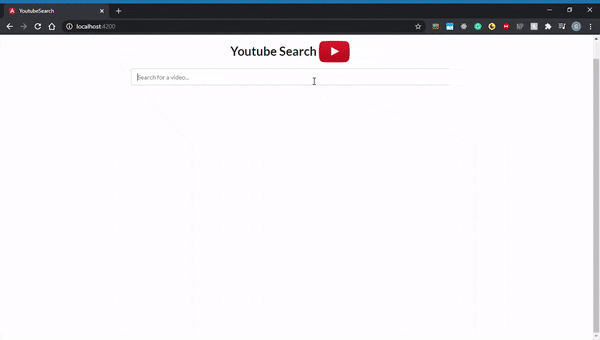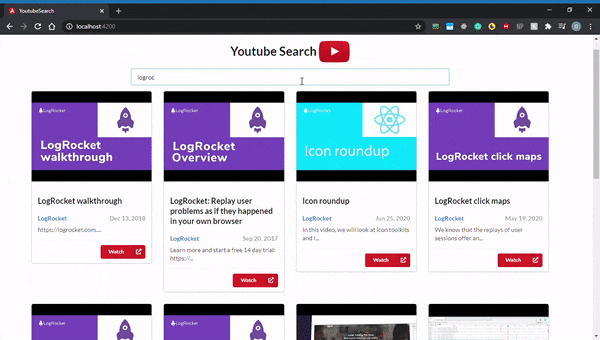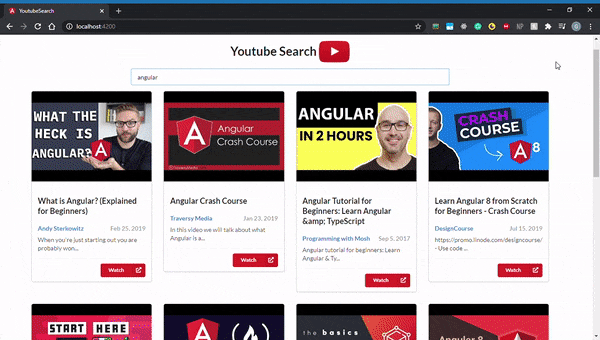
By default, all Angular applications are served at localhost:4200, so go ahead and open that up in your browser. Here’s what it should look like:
Conclusion
You should now have a good understanding of how to use Angular and RxJS to build a YouTube video search application. We walked through how to implement certain core concepts by using them to build a simple application. We also got a sense of RxJS’s powerful features and discussed how it enables us to build certain functionalities with tremendous ease.
Best of all, you got a slick-looking YouTube search app for your troubles. Now you can take the knowledge you gained and implement even more complex features with the YouTube API.
Experience your Angular apps exactly how a user does
Debugging Angular applications can be difficult, especially when users experience issues that are difficult to reproduce. If you're interested in monitoring and tracking Angular state and actions for all of your users in production, try LogRocket.
LogRocket is like a DVR for web apps, recording literally everything that happens on your site including network requests, JavaScript errors, and much more. Instead of guessing why problems happen, you can aggregate and report on what state your application was in when an issue occurred.
The LogRocket NgRx plugin logs Angular state and actions to the LogRocket console, giving you context around what led to an error, and what state the application was in when an issue occurred.
Modernize how you debug your Angular apps – Start monitoring for free.
The post Build a YouTube video search app with Angular and RxJS appeared first on LogRocket Blog.




Top comments (0)