From a comment on the announcement article:
For this contest, you'll only be able to submit projects that you started after this announcement post was published.
Well, I guess I am disqualified. But, leaving this post up as a resource for anyone that may like it.
My Workflow
I am a big fan of Tailscale for a Wireguard VPN mesh. Some months ago, someone requested that Tailscale support an Ansible Galaxy role to set up a Tailscale node. I saw the help wanted tag and knew what I must do.
However, Tailscale supports a large number of operating systems, and I wanted my Tailscale Ansible role to support everything that Tailscale supports. I needed a CI workflow that would test all those systems in parallel whenever I created a new PR.
I still have a few operating systems to add support for (dreading Windows and OSX for the CI difficulties they present), but I currently run tests against 10 operating systems in parallel using a GitHub Action workflow to spawn dynamic Action jobs that each run Molecule against a single system. This leverages a GitHub Action matrix.
Submission Category:
Maintainer Must-Haves
Yaml File or Link to Code
 artis3n
/
ansible-role-tailscale
artis3n
/
ansible-role-tailscale
Ansible role to install and enable a Tailscale node.
artis3n.tailscale
This role initializes a Tailscale node.
Find supported operating systems on this role's Ansible Galaxy page.
Requirements
You must supply a tailscale_auth_key variable, which can be generated under your Tailscale account at https://login.tailscale.com/admin/authkeys.
Role Variables
tailscale_auth_key
Required
An ansible-vault encrypted variable containing a Tailscale Node Authorization auth key.
A Node Authorization auth key can be generated under your Tailscale account at https://login.tailscale.com/admin/authkeys.
Encrypt this variable with the following command:
ansible-vault encrypt_string --vault-id tailscale@.ci-vault-pass '[AUTH KEY VALUE HERE]' --name 'tailscale_auth_key'
See Ansible's documentation for an explanation of the ansible-vault encrypt_string command syntax.
release_stability
Default: stable
Whether to use the Tailscale stable or unstable track.
stable:
Stable releases. If you're not sure which track to use, pick this one.
unstable:
The bleeding edge. Pushed early and often. Expect rough edges!
tailscale_args
Pass any additional command-line arguments to tailscale up.
Note…
The Ansible role can be found on Ansible Galaxy here.
The key for this GitHub Action matrix workflow to work is the following setup in my molecule.yml file:
platforms:
- name: instance
image: ${MOLECULE_DISTRO:-geerlingguy/docker-centos7-ansible:latest}
From there, I can set up which systems to run Molecule against with a matrix:
jobs:
molecule:
runs-on: ubuntu-latest
strategy:
matrix:
distros:
- geerlingguy/docker-centos7-ansible:latest
- geerlingguy/docker-centos8-ansible:latest
- geerlingguy/docker-amazonlinux2-ansible:latest
- geerlingguy/docker-ubuntu2004-ansible:latest
- geerlingguy/docker-ubuntu1804-ansible:latest
- geerlingguy/docker-ubuntu1604-ansible:latest
- geerlingguy/docker-debian10-ansible:latest
- geerlingguy/docker-debian9-ansible:latest
- geerlingguy/docker-fedora31-ansible:latest
- artis3n/docker-arch-ansible:latest
fail-fast: false
steps:
Important! - You don't want to forget fail-fast: false.
This stops the action workflow from stopping whenever a single job fails. We want each spawned job to run in isolation from the others, so we can independently see which systems are good and which have issues.
Then, we just need a single job definition that uses the distro matrix variable. GitHub Actions will automatically spawn a job for each item in the matrix.
- name: Molecule
run: echo "${{ secrets.VAULT_PASS }}" > /home/runner/work/_temp/.vault-ci-pass && ANSIBLE_VAULT_PASSWORD_FILE=/home/runner/work/_temp/.vault-ci-pass pipenv run molecule test
env:
MOLECULE_DISTRO: "${{ matrix.distros }}"
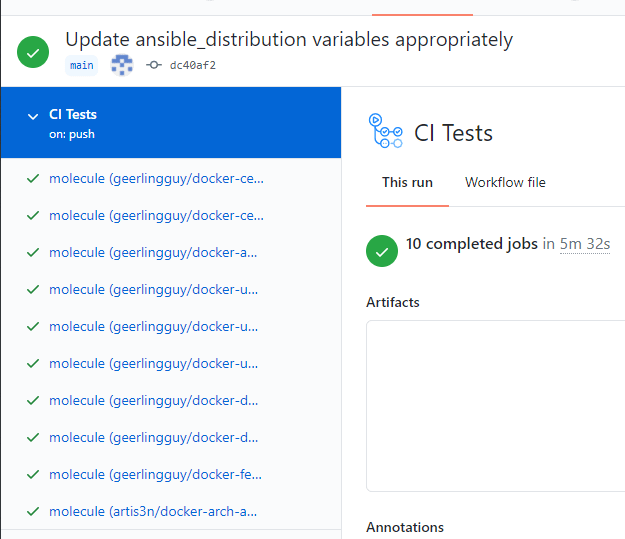
This spawns a job for each matrix distribution:








Top comments (0)