Originally posted on realpythonproject.com
We will be using PuLP and the chemparse libraries to balance chemical equations
You can find the source code here
Pre-Requisites
- Familiarity with Constraint Optimization. Check out my previous article for an introduction to Constraint Optimization and PuLp
- Some Familiarity with Balancing Chemical Equations.
- It is good to have some familiarity with Streamlit.
How can we balance Chemical Equations using Constraint Optimization?
Balancing a chemical equation essentially means respecting the conservation of mass and ensuring the same number of atoms of an element are present on the left-hand side (Reactants) and the right-hand side (Products). Basically if you X grams of an element as a reactant, the product will also have X grams of that element in the Products (ideal conditions).
In the above picture, you can notice the following
- In the unbalanced equations, there are 2 atoms of H and Cl in the Products. However, there is only a single atom of H and Cl in the Reactants. This does not respect the law of conservation of mass
- In the balanced equation, we are using 2 atoms of H and Cl in the reactants. The number of atoms in the Products remains unchanged. As a result, the equation is balanced and it no longer violates the law of conservation of mass.
The above equation was easy to solve and can be done manually. However, things get complicated as the number of elements increases and the number of reactants/products increases. Eg: The below equation although doable manually will require quite a bit of trial and error.

So how can we use Constraint Optimization?
Let’s consider the first equation we had looked at

The coefficient of each reactant/product can be thought of as a variable.

X1, X2, X3, and X4 are the variable coefficients. Each variable must be an integer greater than or equal to 1.
Each element adds a constraint,i.e
- For Zn: The number of Zinc Atoms in Reactant must equal to the number of Zinc Atoms in Products
- For Cl: The number of Chlorine Atoms in Reactant must equal to the number of Chlorine Atoms in Products
- For H: The number of Hydrogen Atoms in Reactant must equal to the number of Hydrogen Atoms in Products
The Problem can be considered as a Minimizing Problem since we need to find the smallest coefficients that can balance the equation.
The problem has no objective other than minimizing the coefficients, which means the objective is basically 0 (None).
To summarize this is how our problem is setup
This problem can be set up using PuLP and solver using PuLP’s default solver.
Now, we can move on to generalizing this to support other equations as well.
Parsing the Chemical Equation
We will use the chemparse library to parse the reactants and products.
pip3 install chemparseBelow are a few examples of what chemparse returns. They are taken from the library’s documentation
“CH4” returns {“C”:1.0, “H”:4.0}
“C1.5O3” returns {“C”:1.5, “O”:3.0}
"(CH3)2(CH2)4" returns {"C":6.0, "H":14.0}Let's create a function to parse the equations. The function will expect an input similar to the one below
Zn + HCL -> ZnCl2 + H2We can split by ‘->’ to get the Left Hand Side and the Right Hande Side]
To get the compounds we can split by “+” and remove the trailing spaces.
We need to store the unique elements so we can form constraints for each and convert each reactant and product into a dictionary produced by chemparse.
First, we iterate over the lhsCompounds and store the result from chemparse as well as the unique elements.
After that we iterate over rhsCompounds, we do not need to store the unique elements again. However, we store the coefficients as a negative number since this will make it easier when setting up our constraints.
Function to set up and solve Constraint Optimization Problem
Now, we can move on to the problem-solving stuff.
We will call the parse function we had written earlier and store the returned values.
Every reactant/product will have a co-efficient, therefore we can simply loop over the variable allCompounds (this contains the output from chemparse for each reactant/product) and create a variable during each iteration.
The variables will have a category of Integer and a lower bound value of 0
As discussed earlier, the problem is a Minimization problem with no Objective.
Now, let’s set up our constraints
As discussed earlier, each element will add a constraint associated with it.
- we loop over the uniqueElements
- get the number of atoms of the element in each reactant/product
- sum up the coefficients (Remember all the Product coefficients are stored as negative values)
- Add a constraint that the sum should be 0
Now, we are ready to solve the problem and create the balanced equation
Streamlit WebApp
Install Streamlit
pip install streamlitThis will be a pretty minimalistic app.
We create a text_input component and set the default value to an unbalanced equation. The balance() function is imported and we pass the text_input’s value as an argument.
Additionally, we can also display the problem before we solve it. This should be inside the balance() function
st.text(prob)




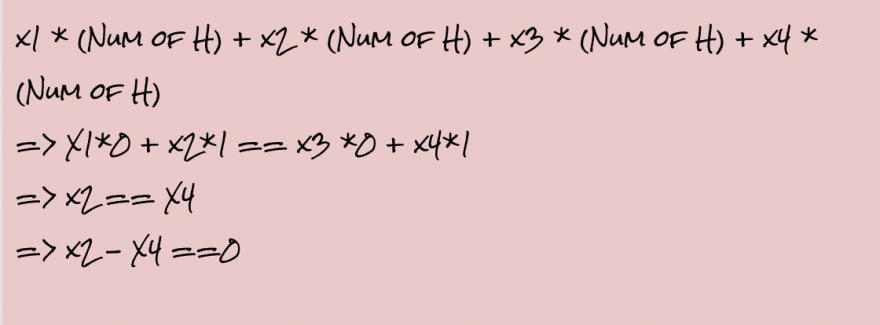









Top comments (1)
I didn't even know that Python could be used to solve chemical equations faster. You wrote that "The above equation was easy to solve and can be done manually", but for me it's definitely not easy. I never understood chemistry well, so I often used uk.edubirdie.com/chemistry-help to somehow complete the necessary tasks and worry less about my success. This allowed me to devote more time and attention to the subjects that interest me, and almost never think about chemistry, because it is good that I have found such a wonderful learning resource. I recommend everyone to try it, you will not regret it!